 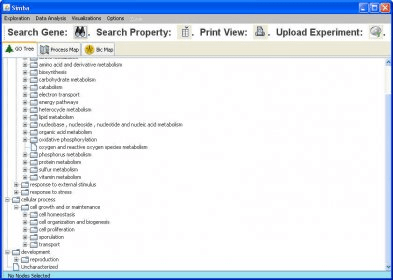
Existing software packages targeted for image-based screening, however, have one or more limitations which prevent sophisticated visualization and extraction of information from image-based screens: (a) they are not designed for the hierarchical data structure inherent in image-based data (each treatment condition is replicated in several samples, each sample is usually represented by several images, each image contains a population of cells, and each cell has hundreds of associated measures), (b) they ignore the inherent biological variability of cell populations such that assays requiring subpopulation analysis cannot be scored, (c) they cannot handle the volumes of data typical in image-based experiments (e.g., ~500 measurements for each of ~100 million individual cells), (d) they provide limited linking to raw or processed image data or chemical structure data, (e) they allow only limited statistical analyses of the data, (f) they are proprietary and new methods cannot be easily added, (g) they are limited to data from a particular image analysis package, (h) they require expertise in statistics or programming, and/or (i) they require intense hands-on data management. 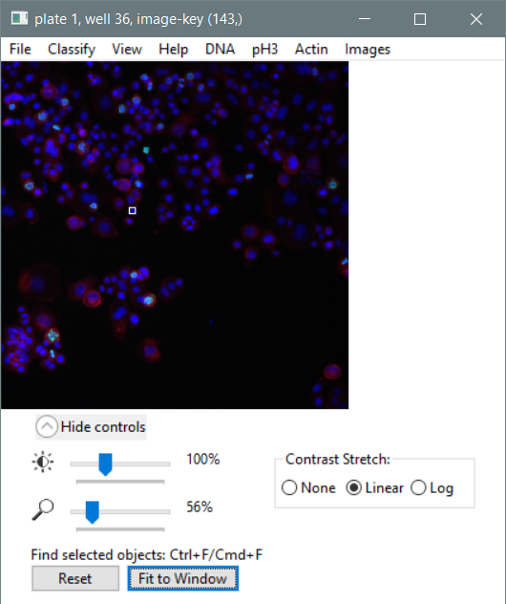
For analysis of small or very simple experiments, spreadsheet programs like Microsoft Excel are sufficient, and useful open-source tools exist for analysis and exploration of data from high throughput screens in general. The volume and richness of individual-cell data from large image-based screens is unprecedented and existing software is inadequate for the challenge of data analysis. This tool has been useful for extracting image-based measurements to score sophisticated screens, with many more in progress. 
We recently developed open-source image analysis software, CellProfiler, which measures a rich set of cellular features in images, such as size, shape, and staining patterns including intensity, texture, and colocalization. This capability is increasingly important given the ease now to systematically perturb cells with libraries of chemicals or gene-perturbing reagents like RNA interference or gene overexpression and collect hundreds of thousands of images of these cell samples. The scientific community has only begun to scratch the surface of computationally extracting the rich information visible in fluorescence microscopy images of cell samples. For this, load the properties file in CellProfilerAnalyst.Visual analysis of cell samples has played a dominant role in the history of biology. A user-friendly option for machine learning is the softwareCellProfiler Analyst. Step 3: Use any programming language for supervised or unsupervised machine learning, such as python or R. The pipeline also generates a CellProfiler Analyst properties file for the machine learning in step 3. The example CellProfiler pipeline exports the features as csv files. Step 2: Segment images and extract features in CellProfiler. The app reads a cif file and writes the tiles (which are tif image files) to the output folder. Step 1: Automatically generate tiles of 1000 single cell images per tile, using a python app (alternatively a Matlab script is available). Preparatory Step: Identify cell populations using gating in IDEAS software. Label-free cell cycle analysis for high-throughput imaging flow cytometry. An open-source solution for advanced imaging flow cytometry data analysis using machine learning. (2016), however, the former protocol is still available here. Note: This is a more user-friendly and streamlined protocol as compared to Blasi et al. This high-dimensional data can then be analyzed using cutting-edge machine learning and clustering approaches using user-friendly platforms such as CellProfiler Analyst or scripting languages such as R or Python. The image tiles are imported into the open-source software CellProfiler, where an image processing pipeline identifies cells and subcellular compartments allowing hundreds of morphological features to be measured. 
cif file format) can be read and resulting image tiles are generated. Compensated data files from an imaging flow cytometer (the proprietary. It will help to reveal otherwise unappreciated populations of cells based on features that may be hidden to the human eye. This protocol aims to enable the scientific community to leverage the full analytical power of IFC-derived data sets. We here provide an open-source IFC protocol described in Hennig et al. CellProfiler can be used to analyze the resulting images from imaging flow cytometry, whether brightfield, darkfield, or fluorescence. Imaging flow cytometry (IFC) combines the high-throughput capabilities of conventional flow cytometry with single-cell imaging. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
